The Roadmap to Fight “Cantonese” Cancer
The pioneering position in genetic and genomic research on nasopharyngeal cancer (“NPC”) at CUHK was anchored by the late Prof. Dolly P. Huang (1935–2004). Using various genome-wide approaches such as allelotyping, comparative genomic hybridization (“CGH”) and microarray analysis, Professor Huang’s group was the first to report the following findings:
- key genetic changes in NPC patients as indicated by the loss of short arm of Chromosome 3
- alterations in the p16 and RASSF1A genes drive the development of NPC
- NPC genome map
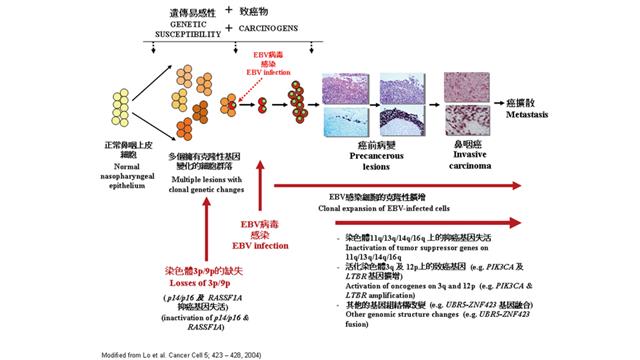
Professor Huang’s protégé, Prof. Lo Kwok-wai of the Department of Anatomical and Cellular Pathology, continues the legacy and contributes significantly to the research in NPC. The NPC tumorigenesis model they published in Cancer Cells in 2004 has since been well recognized in the medical circles.
Why study Nasopharyngeal Cancer?
NPC is a common cancer in Asia. 80% of the cases occur in Southern China and among Chinese in the Southeast Asian region, a much higher percentage than their western counterparts. Scientists suggested that a combination of environmental factors, like dietary preference, infection by the Epstein-Barr virus (EBV) and genetic make-up could result in the onset of NPC. For the past two decades, Professor Lo’s group found that the subsequent genetic and epigenetic abnormalities after the EBV infection are necessary to drive the tumorigenic process.
At its early stages, NPC can be effectively treated by radiotherapy. Unfortunately, the symptoms of NPC are highly nonspecific until it is in an advanced stage, when treatment is much more difficult and less effective. Professor Lo’s group is engaged in the quest to identify the oncogene of NPC to enable efficient treatment of NPC as well as development of molecular target drug.
Recent discoveries of genetic and epigenetic changes in NPC
Professor Lo and his collaborators commenced a systematic screening for genetic abnormalities involved in the development of NPC. They have studied a panel of tumour tissue samples, cell lines and xenografts. The array-based CGH process they used to map the genetic information of the NPC tumours provided a number of possible NPC oncogene targets for further research in NPC. Specifically, they have performed functional studies on the CCND1 gene at chromosome 11q13, which encodes the protein Cyclin D1 and the LTBR gene at chromosome 12p13, which encodes the protein lymphotoxin beta receptor. They have found that the amplification and over expression of these proteins are common in the primary NPC tumour samples. Both of them were shown to be critical to the development of NPC tumours.
Another international collaboration between local colleagues and those from Canada, France and Korea discovered a novel UBR5-ZNF423 transforming fusion gene in 8.3% of NPCs by whole-transcriptome sequencing. The expression of this gene might contribute to the transformation of a subset of NPCs. This is the first evidence to support the important role of gene rearrangement in NPC.
Next milestone, the finest genome map of NPC
These important findings lay the foundation for the development of targeted therapy for NPC by inhibiting the NPC-associated oncogenes such as the UBR5-ZNF423 protein. Using massive parallel genome sequencing, the group plans to catalogue the whole spectrum of genomic changes involved in NPC tumorigenesis. The findings from this endeavour will provide important novel biomarkers and therapeutic targets for the development of personalized cancer therapy.
The Basics